Kitzerow’s Jigsaw Puzzle Methodology
The Jigsaw Puzzle Research Methodology
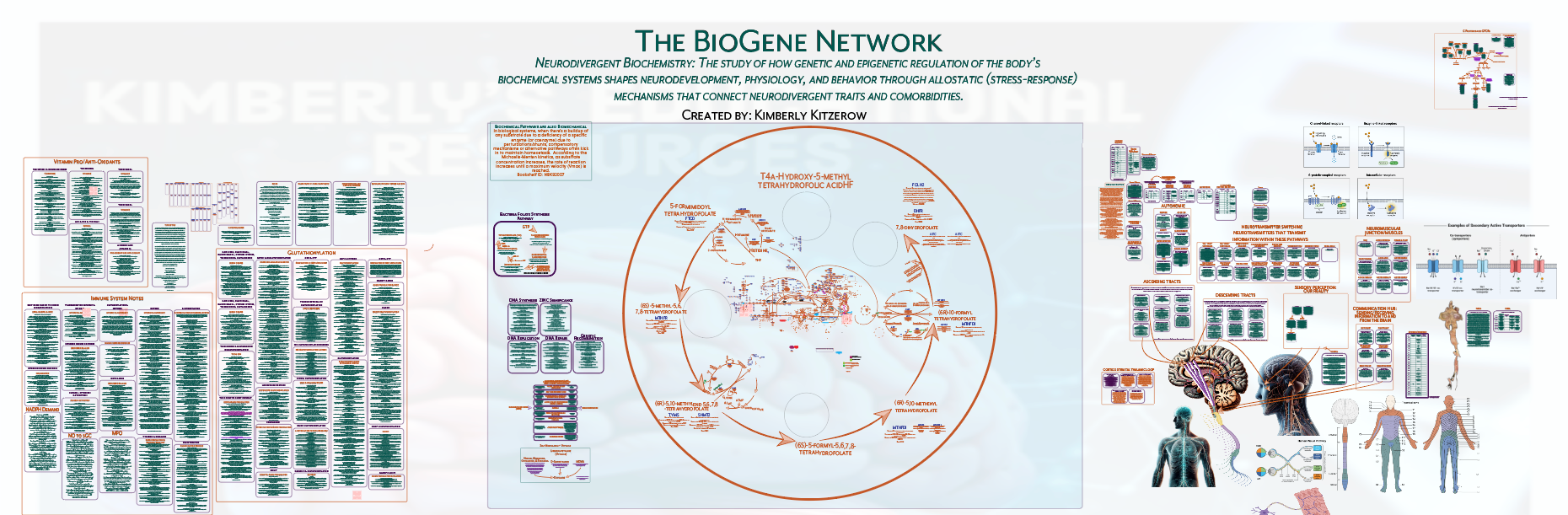
A computational systems-analysis methodology in which a biochemical network of gene-coded proteins is constructed to define a conserved species-level functional framework, then used as a reference structure to compare demographic-level datasets against the broader population and species-level architecture.
The methodology treats complex biology as an integrated system rather than isolated findings. Biomarkers, pathways, and studies are evaluated as interconnected components of a cohesive biological structure.
A species-level biochemical network of gene-coded proteins establishes the conserved reference architecture against which demographic-level autism and comorbidity patterns can be systematically compared.
Demographic-level datasets are compared against the species-level biochemical reference network to identify biologically constrained points of dysregulation that distinguish the specific demographic from the conserved species-level framework.
The biologically constrained points of dysregulation identified through this species-to-demographic comparison were organized into the resulting theoretical cascade. Public ResearchGate papers are provided as supporting literature review and converging evidence demonstrating alignment across independently published findings, biomarker datasets, and systems-level biological relationships.
Scientific Method and Method Application
This work follows the broader structure of the scientific method, from observation and research through analysis, interpretation, communication, and public education. The Jigsaw Puzzle Research Methodology operates within the testing and analysis phases by comparing a conserved species-level framework to demographic-level biomarker data in order to identify biological mechanism.
The process begins with the repeated co-occurrence of autism traits and comorbidities, indicating that these outcomes follow a structured biological pattern rather than appearing randomly.
Existing autism research, biomarker findings, and protein-level biological data were reviewed to determine whether independent lines of evidence converged on shared systems, pathways, or regulatory disruptions.
The Exclusivity Principle was proposed: autism traits and comorbidities are not independent conditions but emerge together from a shared underlying biological mechanism. Their co-occurrence reflects a unified biochemical origin rather than separate, unrelated causes.
The Jigsaw Puzzle Research Methodology was applied as a computational systems analysis method. A biochemical network of gene-coded proteins was constructed to define a conserved species-level functional blueprint and then used as a reference framework for comparison against demographic-level biomarker datasets.
A biochemical network of gene-coded proteins was constructed to represent the conserved biological architecture used as the reference system.
Autism-associated biomarker datasets were mapped onto this network to align population-level data with the species-level framework.
Deviations from the reference framework were identified as consistent points of dysregulation across datasets.
Recurrent patterns across pathways and regulatory systems were traced to reconstruct the biochemical cascade associated with the observed traits and comorbidities.
Identified points of dysregulation were analyzed for convergence to determine whether the same pathway-level disturbances appeared consistently across independent datasets.
Based on repeated convergence, a coherent biochemical cascade was reconstructed linking autism traits and comorbidities to shared system-level dysregulation, supporting the Exclusivity Principle.
The framework was documented and shared publicly so that the underlying logic, evidence structure, and biological interpretation could be reviewed, scrutinized, and compared against future findings.
Watch the documentary → View the documented timeline →Educational frameworks such as Neurodivergent Biochemistry, BioToggles, and BioDials were developed to translate system-level mechanisms into structures that can be understood, evaluated, and applied across audiences.
Learn more about the core frameworks →Testable Pillars of Kitzerow’s Autism and the Comorbidities Theoretical Model
This model can be evaluated at defined system-level stages where regulatory activation, pathway behavior, neural circuitry, and downstream outcomes converge. These pillars are integration points built on mechanisms already studied at the micro level, allowing the cascade to be tested through how those mechanisms interact across systems and over time.
View Research Papers Outlining the Mechanisms →Stress Activation
Genetic and epigenetic factors activate internal stress-response systems across regulatory domains, including the immune system, metabolism, cellular repair, nervous system, and genetic regulation.
These activations may be situational, chronic, or genetically driven. The duration and category determine downstream biological effects.
Do genetic and epigenetic mutations produce a convergent and sustained stress-response state across regulatory systems?
BH4 Pathway Shunt
Stress-response activation redirects biochemical pathway activity through the redox-regulated, GCH1-mediated BH4 Shunt, shifting activity across AAAH, NOS, and AGMO pathways.
This coordinated redistribution links multiple physiological systems and creates shared biochemical conditions underlying both autism and comorbid traits.
Does stress-induced BH4 pathway redirection produce biochemically linked autism and comorbid trait clustering?
Neural Circuit Disruption
The AAAH pathway shifts aromatic amino acids away from monoamine synthesis and toward glutamate production, altering neurotransmitter balance.
This contributes to excitatory and inhibitory imbalance within cortico-striatal-thalamic circuitry, which drives the expression of autism traits.
Does disruption of excitatory and inhibitory balance within CSTL circuitry produce autism traits?
Comorbidity Clustering
NOS Shunt-induced epigenetic redox-sensitive protein shunts function as regulatory effectors that alter biochemical pathway activity across systems.
These shifts disrupt coordination across biological timing cycles and produce consistent clustering of autism traits and comorbid conditions over time.
Do genetic and epigenetic factors alter biochemical pathway activity, producing consistent clustering of autism and comorbid traits?
How the Research Maps to the Framework
Each tested framework component is presented as a question, followed by research that directly evaluates it.
Stress Activation
2025 Japanese Study
BH4 Pathway Shunt
2025 Brazilian Study
Neural Circuit Disruption
Stanford + Yale Studies
Comorbidity Clustering
2025 Princeton Study
Explore the full breakdown of how each mechanism has been tested across independent studies and how the cascade aligns at the system level.
Methodology Questions and Clarifications
Why does the method use a species-level baseline?
The method uses a species-level baseline because it is based on the function of proteins. Protein function is conserved at the species level, which provides a stable biological reference across pathways and regulatory systems.
What the Baseline Represents
A reference system defined by conserved protein functions across biological pathways and domains.
What Is Conserved
The functional role proteins serve within biological systems remains consistent at the species level.
What Is Not Assumed
The method does not assume identical expression, identical biomarkers, or identical phenotypes across individuals.
Why This Matters
A stable functional baseline is required to determine whether observed biological patterns reflect expected variation or structured system-level deviation.
The baseline is species-level because protein function is species-level. That conserved function defines the reference used to evaluate biological variation.
Where does individual variation enter the method?
Individual variation is determined by genetic and epigenetic factors that influence protein function. These factors affect how proteins operate within biological systems and how strongly those effects are expressed at the pathway and system level.
Source of Variation
Genetic and epigenetic factors alter protein behavior, influencing how biological systems respond and regulate.
What Changes
Variation occurs in protein efficacy and in the downstream impact those changes have across pathways and regulatory domains.
What Biomarkers Reflect
Biomarkers reflect the observable outcomes of these differences in protein function across biological systems.
What Is Evaluated
The method evaluates whether these differences form recurring system-level patterns rather than isolated variation.
Individual variation reflects how genetic and epigenetic factors influence protein function and system behavior, not variation in whether proteins have function.
What is the comparative layer of the method?
The comparative layer evaluates how observed biological states relate to conserved protein function across systems. It determines whether biomarker patterns reflect expected variation or consistent system-level deviation.
Reference System
A species-level baseline defined by conserved protein function.
Observed Data
Population-level biomarker datasets representing measured biological states.
Evaluation Focus
Whether observed patterns align with expected biological behavior or reflect structured deviation.
Output
Recurring system-level patterns that can be evaluated for biological consistency.
The comparative layer evaluates biological alignment at the system level. It does not disclose the internal structure used to generate that evaluation.
What makes a result meaningful within the method?
A result is meaningful when it is not isolated, when it recurs across datasets, and when it aligns with the broader organization of biological systems.
Recurrence
The same type of deviation must appear across datasets rather than as a single isolated finding.
Coherence
Observed deviations must fit within pathway structure and regulatory relationships rather than stand alone.
System Alignment
Patterns must align across biological domains in a way that is consistent with the larger model structure.
Interpretive Threshold
Only convergent patterns are treated as sufficient for reconstructing the biochemical cascade.
A meaningful result is not just a difference. It is a recurring difference that aligns across system structure and biological function.
Why is part of the method protected as a trade secret?
The internal system architecture underlying the model is protected as a trade secret under IP attorney guidance. This decision followed uncited use of the model and lack of institutional correction.
What Is Public
The structure of the method, its evaluation logic, and its system-level outputs.
What Is Protected
The underlying system architecture and implementation framework used to generate the model.
Why It Is Protected
Protection was implemented under legal guidance following uncited use and lack of correction.
What Can Be Evaluated
The model can be evaluated through its structure, predictions, and alignment with independent research.
The method is transparent in what it evaluates and produces, while protecting the underlying system that generates those outputs.

