
My Process - My How and My Why
Who Am I and Why Did I Do This?
I am an educator and neurodivergence advocate who collaborates with educators, researchers, and clinicians to increase understanding of neurodivergence and its associated comorbidities. My work began in October 2020, when my nonverbal autistic daughter could not blow out the candles on her fourth birthday cake. That moment shifted my view of her nonverbal autism from a behavioral interpretation to a physiological one, and I recognized that nonverbality was a comorbidity rather than part of autism itself. I also realized that many comorbidities in autism are physiological, which raised important questions about when high prevalence signals an underlying mechanism within the condition.
This insight led me to create NeuroToggle, an applied neuroplasticity framework designed to support the development of all neural circuits, both motor and non motor, because every neural circuit is a behavior or skill that can be intentionally targeted through neuroplasticity instruction strategies that utilize established teaching pedagogy.
As my daughter progressed, and I posted our story on social media, people asked how her nonverbal state related to her autism diagnosis and why autistic individuals often experience overlapping medical conditions. To answer these questions, I began mapping biomarkers and biochemical pathways to identify the mechanisms that link autism traits with their comorbidities. This work resulted in the Autism and the Comorbidities Theory and the broader systems level framework I call Neurodivergent Biochemistry. I document this research through books, ResearchGate papers, visual frameworks, and a timestamped timeline to support scientific clarity and cross disciplinary collaboration.
-
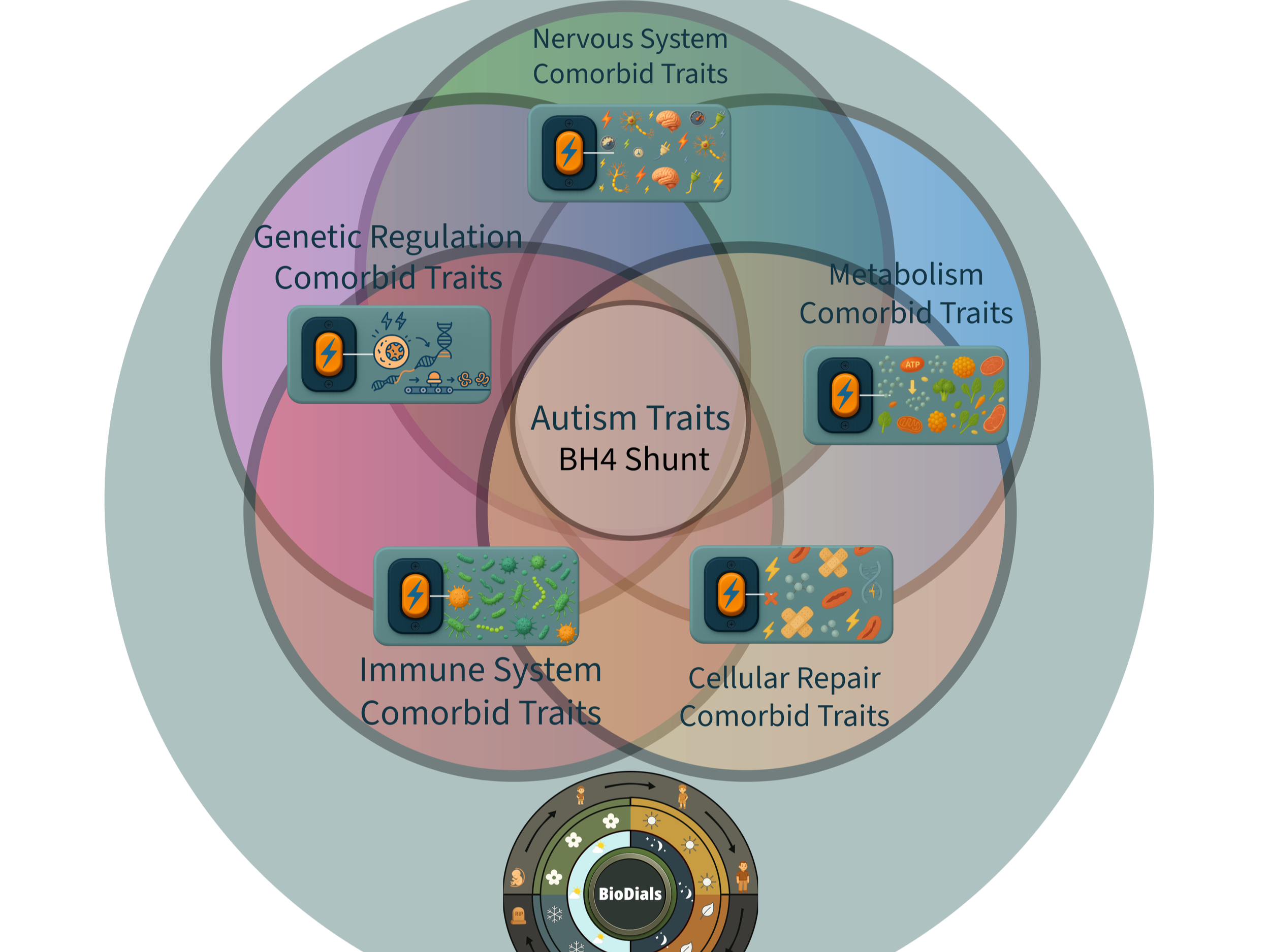
Why I Went For the Link Between Autism and the Comorbidities
Following the public documentation of my daughter’s progress, specifically her achievement of functional speech, I was frequently asked how her nonverbal state related to her autism diagnosis. To explore this question, I began visually mapping potential connections using Adobe Illustrator, arranging peer-reviewed data into a jigsaw puzzle format to identify cross-disciplinary patterns.
This method allowed me to systematically examine overlapping biochemical, neurological, and immunological mechanisms. Through this process, I observed that biomarker data consistently pointed to dysregulation within a specific biochemical pathway. More precisely, the data revealed that this pathway was not simply underactive or overactive, but shunted—rerouted in a way that reflected a physiological shift in priorities under chronic stress.
The recurring presence of both autism traits and systemic comorbidities (At rates from 50-80% prevalence per comorbid condition, and more than 95% of autistics having at least one of them) in a specific subset of individuals suggested a unified, simultaneous genetic root cause. Comorbidities include sleep disorders, GI problems, metabolism disorders, neuroinflammation, altered inflammatory responses, immune abnormalities, anxiety and other mental health conditions, neurological disorders, lysomal storage disorders, seizures, tics, OCD, ADHD, motor delay, motor impairment, obesity, deafness, hearing loss, Functional Neurological Disorder (FND), Hypermobile Ehlers-Danlos syndrome (hE-DS), postural tachycardia syndrome (PoTS) and learning differences (Al-Betagi, 2021; Chung & Kim, 2024; Khachadourian et al., 2023; Bougeard et al., 2021; Burns et al., 2023; Hours et al., 2022; González-Herrero et al., 2022; Owens et al., 2021). This led to the identification of converging disruptions centered on the tetrahydrobiopterin (BH4) pathway, a pattern I would later define as the BH4 shunt.
I soon observed that this shift may also involved redox-sensitive proteins, which are proteins that react to reactive oxygen species, etc. to restore redox homeostasis. Redox homeostasis is maintenance of the balance between antioxidants vs antioxidants, the balance of which is obtained by redox regulation which involves redox sensing, signaling, responses, and feedback control pathways. This is part of our regulatory system and reduction of oxidative modification is fed by NADPH and NADH. Cellular protein thiols are the primary targets of redox modification and this is mediated by the GSH (thioredoxin and glutathione) systems (Sies et al., 2024) . When such restoration fails, a progressive state of allostatic load ensues. This load may suspend the proteins in a state of allostatic regulation—away from baseline conditions that support the participation of the proteins that are required for typical neurodevelopment and physical growth. More research needs to be done in this area as to which developmental proteins are disrupted via stress mechanisms, and how that impacts physical and neural development.
This realization became the foundation for what would ultimately evolve into my integrated framework: BioToggles, the biochemical network map, and the overarching model of Neurodivergent Biochemistry, a system in which redox shifts and protein isoform selection mediate systemic adaptations that manifest as both behavioral and physiological phenotypes. Existing in a state of allostasis vs baseline homeostasis.,
-
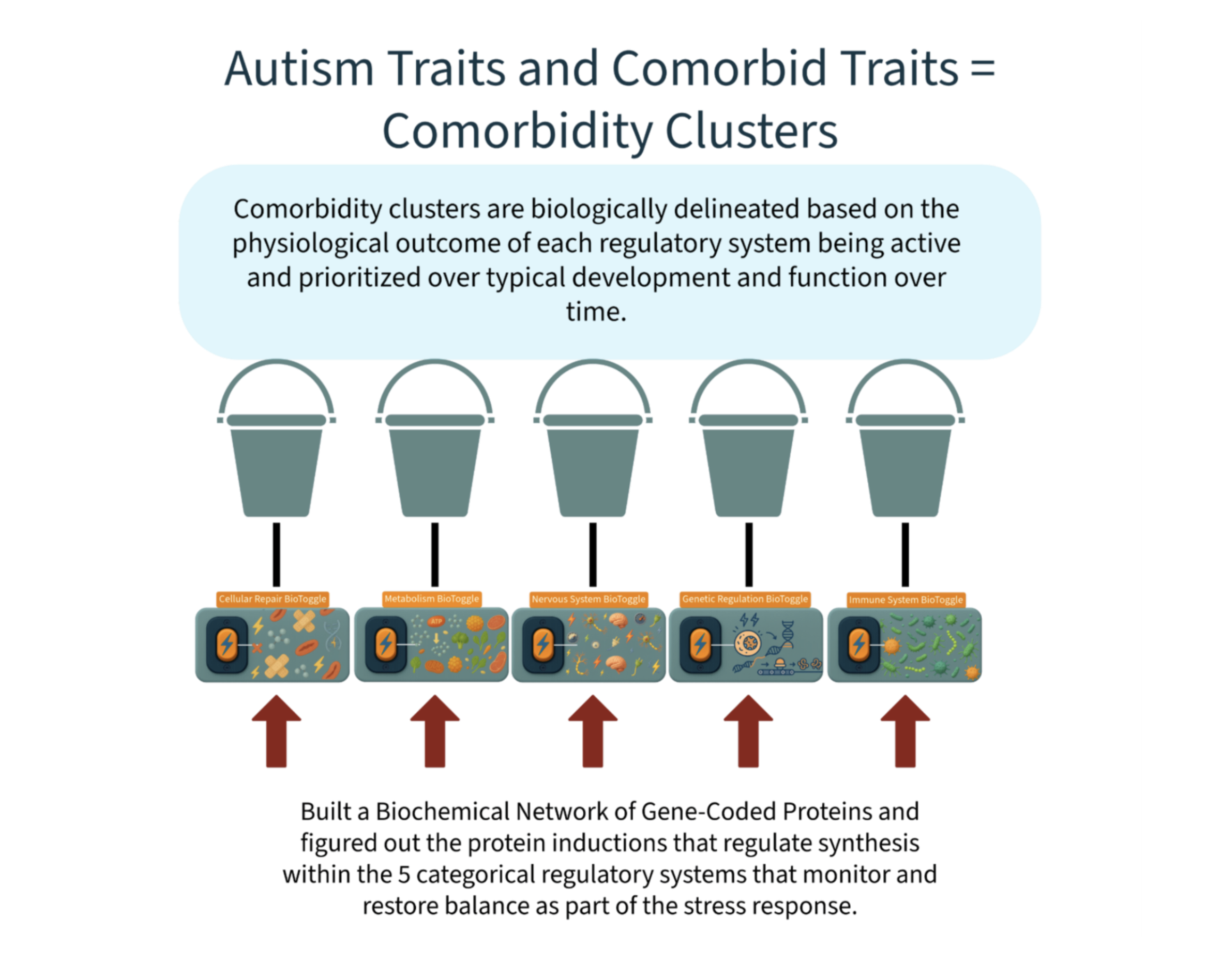

Why I Went Looking For the Answer In Our Biochemistry
Biochemistry stands apart from interpretive sciences because it is rooted in observable, reproducible processes governed by universal chemical and physical laws. Biochemical pathways (ranging from the TCA cycle and glucose metabolism to complex signaling cascades and neurotransmitter synthesis) operate with predictable precision, utilizing the same proteins and cofactors for each reaction across all humans.
Over 99.9% of protein-coding genes are conserved across individuals (Collins & Mansoura, 2001). However, due to genetic variation, including single nucleotide polymorphisms (SNPs), alternative splicing, and post-translational modifications , subtle differences can emerge in isoform expression or protein activity. Still, the core instructions for protein synthesis are encoded in DNA and can be directly measured and validated.
This inherent objectivity distinguishes biochemistry from fields that rely on statistical correlations or associative studies, which often depend on interpretive frameworks and are subject to revision as new data emerge. By utilizing metabolomics (Johnson et al., 2016), and comparing biochemical pathways with metabolite biomarker data in individuals who a given condition or diagnosis, researchers can establish a mechanistic understanding of how those pathways are functioning in those who have obtained that diagnosis—and how such disruptions influence the biochemical system as a whole. These comparisons enable precise identification of molecular dysfunctions underlying complex conditions, offering clarity where correlation-based approaches fall short. This becomes even more useful in individual analysis in what the physiological outcome of biochemical disruption would be for an individual.
-

Why I Created a Biochemical Network and the Jigsaw Puzzle Methodology
I created a biochemical network because I needed to understand the biochemical context behind the metabolite dysregulation seen across autism and its comorbidities. The biomarker data were consistently abnormal, and I wanted to know what that pattern meant at the level of protein function. To answer this, I pulled raw protein data from UniProt and built a large scale biochemical network in Adobe Illustrator so I could visualize how proteins, pathways, metabolites, and redox regulated processes interact across systems.
Once the network was constructed, it gave me a wide scope view of why certain biomarkers were dysregulated at specific points and how protein inductions involving redox contributed to those shifts. It showed how stress responsive pathways interconnected and how biochemical load moved across systems in predictable ways. With the network in front of me, I could see patterns that were not visible when the data remained isolated in separate studies.
To organize all of this information, I formalized a process I call the Jigsaw Puzzle Methodology. This method involves placing peer reviewed findings and protein data side by side, similar to how pieces of a puzzle are arranged until the overall image becomes clear. By aligning shared biomarkers, pathway overlaps, and redox driven protein shifts across neuroscience, immunology, metabolism, endocrinology, and molecular genetics, the underlying mechanism began to emerge.
Constructing this network and applying the Jigsaw Puzzle Methodology allowed me to identify a consistent biochemical pattern that linked autism traits with their comorbidities. This integrative view revealed a genetically induced allostatic state that produced epigenetic and redox sensitive protein shunts, with the BH4 Shunt functioning as the central linchpin of the stress response system. This ultimately led to the development of the Autism and the Comorbidities Theory and the broader framework I now call Neurodivergent Biochemistry.

